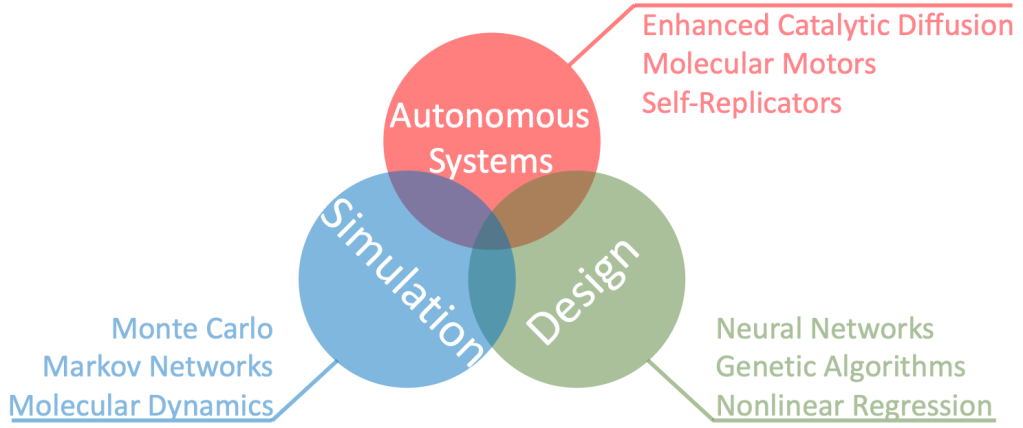
My long-term goal is to create Computer-Aided Design for Autonomous Chemical Systems. This suite of computational methods would allow me to design and optimize systems like molecular motors and self-replicators for a desired function. I hope to one day see artificial machines of my own design working alongside our bodies’ natural processes to heal and repair us. This grand ambition will begin with the three more modest projects discussed below.
Project 1: Understanding Biological Motors
“Thermodynamics owes more to the steam engine than the steam engine to thermodynamics.”
Lawrence Joseph Henderson
Molecular motors drive life processes like muscle contraction, molecular transport, and cell communication. While the general principles of molecular motor operation are known, many details are not. To better understand these machines I propose developing and simulating reactive, coarse-grained models. Biological motors are driven by reactions like ATP hydrolysis. To simulate motors I will develop classical reactive models for ATP (above, top left). The reactivity is modeled with a many-body potential that reproduces a reaction energy profile (above, top right). Models of the motor itself will need to be coarse-grained and parameterized in order to observe complete dynamics (above, bottom left). By creating a nonequilibrium steady state where ATP is injected and hydrolysis products ADP and Pi are removed, I can study the complete dynamics of the motor (above, bottom right).
Project 2: Designing for Dynamics
“What I cannot create, I do not understand.”
Richard Feynman
The most important direction of my proposed work is to design chemically-fueled autonomous systems for desired function. Using models developed in Projects 1 & 3, nonequilibrium simulation will be combined with a variety of tools that will design and optimize these systems. Genetic algorithms (above, top left) will be used to design molecular motors for a desired function, for example. The desired function optimizes over many generations of the algorithm (above, top right). I will augment this approach with a neural net (above, bottom left). I will also use the neural net independently to target a desired function by optimizing input through the backpropagation algorithm. Nonlinear regression (above, bottom right) is also an attractive candidate, as it creates a natural connection between dynamic forces (function) and simple force field terms (design), with the gray inset showing an example three particle structure and how its structure and forces connect to the mathematical structure of the regression.
Project 3: Models of Living Molecules
While an exact definition of life is hard to pin down, life is often characterized by functions like energy conversion, reproduction, organization, growth, and regulation. I would like to study these processes at the molecular level. The molecular motors of Project 1 exhibit energy conversion, taking chemical energy and converting it into mechanical motion. Do catalysts exhibit this kind of behavior? Recent experiments show that catalysts and enzymes diffuse more in the presence of their substrates. By using nonequilibrium simulation (above, left) I will answer the question of whether these molecules are transducing chemical energy to work like living systems and what the mechanism is.
Building systems that reproduce is an important step on the road to creating artificial life. But in order to achieve Darwinian evolution a self-replicating system must achieve exponential growth, a difficult task. Modeling and simulating self-replicating systems will allow the study of the thermodynamics, kinetics, and optimization of self-replication. I hope to be able to illuminate design principles for creating self-replicators that achieve exponential growth. One possible system to start this study is RNA (above, middle), which has self-replicating properties (above, right).